CIRCOS圈图绘制 - 最简单绘图和解释介绍了CIRCOS的安装、基本的配置文件的解释、如何最简单的获得一个CIRCOS图。最主要的部分还是配置文件的位置信息和各个参数的含义解释。
本篇则处理染色体层面展示时用到的配置参数,若有困惑的请先参考上一篇。如果两篇都没有讲明白,请留言。
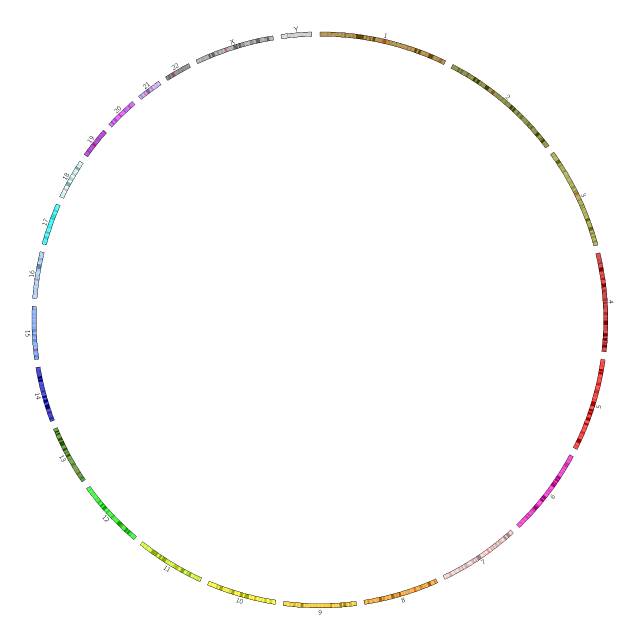
展示染色体染色条带数据
把前面的配置文件再拓展一些,给染色体加上名字,并且按照染色深浅上色。
karyotype变量指定了绘制CIRCOS图所必须的一个文件 (文件的内容虽然通常是染色体的信息,但不局限于染色体信息,其它的区域信息、时间序列信息都可以使用)
文件内容如下 (#后面是注释,会被忽略)
前两列是固定的,chr -。chr表示定义一条染色体;-表示指定这个区域的父区域,染色体没有父区域,用-代替。
ID是当前区域的名字,其子区域的父区域列都使用这个名字。如果同时绘制多个物种,可在ID中包含物种的代号。
LABEL是当前区域显示的名字。
START END是当前区域的范围,必须是整个的区域。如果想显示部分区域,可在后续配置中修改。
COLOR是当前区域的颜色,CIRCOS为每个染色体有定义好的颜色,存储于etc/colors.conf。除了预先定义了染色体的颜色,还定义了一些颜色变量可以直接使用。
- #chr - ID LABEL START END COLOR
- chr - hs1 1 0 248956422 chr1
- chr - hs2 2 0 242193529 chr2
- chr - hs3 3 0 198295559 chr3
- chr - hs4 4 0 190214555 chr4
- chr - hs5 5 0 181538259 chr5
- chr - hs6 6 0 170805979 chr6
- chr - hs7 7 0 159345973 chr7
- chr - hs8 8 0 145138636 chr8
- chr - hs9 9 0 138394717 chr9
如果有cytogenetic band信息,则写到文件末尾。band是固定的,DOMAIN指定从属信息,对应于上面染色体的ID信息。其它列与上面解释相同。
- #band DOAMIN ID LABEL START END COLOR
- band hs1 p36.33 p36.33 0 2300000 gneg
- band hs1 p36.32 p36.32 2300000 5400000 gpos25
- band hs1 p36.31 p36.31 5400000 7200000 gneg
- band hs1 p36.23 p36.23 7200000 9200000 gpos25
- band hs1 p36.22 p36.22 9200000 12700000 gneg
- band hs1 p36.21 p36.21 12700000 16200000 gpos50
- # karyotype定义染色体的名字、ID、起始位置信息,是绘制图的根本
- karyotype = data/karyotype/karyotype.human.txt
- # 必须的部分,控制染色体信息显示
- # http://circos.ca/documentation/tutorials/reference/ideogram/
- <ideogram>
- # 定义显示的染色体之间的间距,为图形半径的0.5% (r代表radius,半径)
- <spacing>
- default = 0.005r
- </spacing>
- # 染色体区域的绘制位置,默认所有染色体都处于远离圆心同样距离的位置
- # 这里设置的是图形半径的0.9倍的位置
- # 也可以设置绝对像素值
- radius = 0.9r
- # 染色体区域的宽度,可以是相对图形半径,也可以说绝对像素值
- thickness = 20p
- # 染色体区域填充颜色
- fill = yes
- # 染色体边的颜色和宽度
- stroke_thickness = 2
- stroke_color = black
- # 显示染色体标签名字
- # 更多label的调整见
- # http://circos.ca/documentation/tutorials/ideograms/labels/lesson
- show_label = yes
- label_font = default
- # ideogram外圈再加总半径的0.05
- label_radius = dims(ideogram,radius_outer) + 0.005r
- # 如果想把label标记在圈里可以写,外圈半径+内圈半径除以2
- # label_rdius = (dims(ideogram,radius_outer)+dims(ideogram,radius_inner))/2
- label_size = 24
- # label与圈平行
- label_parallel = yes
- label_case = upper
- # 显示karyotype中定义的细胞遗传学条带
- # fill_bands=yes 表示使用预先定义的颜色
- show_bands = yes
- fill_bands = yes
- band_transparency = 4
- </ideogram>
- ################################################################
- # The remaining content is standard and required. It is imported
- # from default files in the Circos distribution.
- #
- # These should be present in every Circos configuration file and
- # overridden as required. To see the content of these files,
- # look in etc/ in the Circos distribution.
- # 这些都是引用文件,暂时不去管什么意思,后面用到再逐个解释。
- # 但是绘图时这些必须引用。下面会解释下最关键的引用位置。
- <image>
- # Included from Circos distribution.
- <<include etc/image.conf>>
- file* = circos_label_band.png
- </image>
- # RGB/HSV color definitions, color lists, location of fonts, fill patterns.
- # Included from Circos distribution.
- <<include etc/colors_fonts_patterns.conf>>
- # Debugging, I/O an dother system parameters
- # Included from Circos distribution.
- <<include etc/housekeeping.conf>>
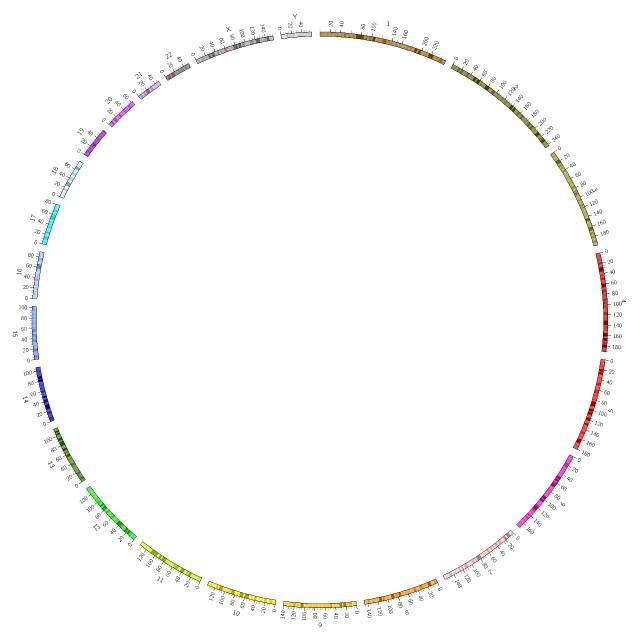
展示染色体刻度信息
显示染色体刻度信息,chromosome_units定义染色体一个单位的大小,缩写为u。若chromosome_units=1000000, 则10u=10000000。
- # karyotype定义染色体的名字、ID、起始位置信息,是绘制图的根本
- karyotype = data/karyotype/karyotype.human.txt
- # 必须的部分,控制染色体信息显示
- # 更多解释见
- # http://circos.ca/documentation/tutorials/reference/ideogram/
- <ideogram>
- # 定义显示的染色体之间的间距,为图形半径的0.5% (r代表radius,半径)
- <spacing>
- default = 0.005r
- </spacing>
- # 染色体区域的绘制位置,默认所有染色体都处于远离圆心同样距离的位置
- # 这里设置的是图形半径的0.9倍的位置
- # 也可以设置绝对像素值
- radius = 0.9r
- # 染色体区域的宽度,可以是相对图形半径,也可以说绝对像素值
- thickness = 20p
- # 染色体区域填充颜色
- fill = yes
- # 染色体边的颜色和宽度
- stroke_thickness = 2
- stroke_color = black
- # 显示染色体标签名字
- # 更多label的调整见
- # http://circos.ca/documentation/tutorials/ideograms/labels/lesson
- show_label = yes
- label_font = default
- # ideogram外圈再加总半径的0.05
- label_radius = dims(ideogram,radius_outer) + 0.05r
- # 如果想把label标记在圈里可以写,外圈半径+内圈半径除以2
- # label_rdius = (dims(ideogram,radius_outer)+dims(ideogram,radius_inner))/2
- label_size = 24
- # label与圈平行
- label_parallel = yes
- label_case = upper
- # 显示karyotype中定义的细胞遗传学条带
- # fill_bands=yes 表示使用预先定义的颜色
- show_bands = yes
- fill_bands = yes
- band_transparency = 4
- </ideogram>
- chromosomes_units = 1000000
- show_ticks = yes
- show_tick_labels = yes
- <ticks>
- # ticks块定义了全局水平的tick的属性
- # 更多解释见http://circos.ca/documentation/tutorials/ticks_and_labels/basics/
- # ticks出现的位置
- radius = dims(ideogram,radius_outer)
- # The label multiplier is the constant used to multiply the tick value to obtain the tick label. For example, if the multiplier is 1e-6, then the tick mark at position 10, 000, 000 will have a label of 10. The multiplier is applied to the raw tick value, regardless of the value of chromosomes_unit.
- # multiplier是用于获得ticks label的,当前ticks对应的染色体位置乘以multiplier就得到ticks label
- # 这个值的设置与chromosomes_units是没有任何关系的
- # 比如当前位置是10,000,000, 乘以multiplier (1e-6)就是10
- multiplier = 1e-6
- # ticks颜色、粗细、大小
- color = black
- thickness = 2p
- size = 15p
- # 默认所有染色体都显示, 放置在ticks块中全局有效
- chromosomes_display_default=yes
- # 不在任何染色体显示, 放置在tick块中单个tick有效
- # 单词拼错无效
- #chromosomes_display_default=no
- # 每个tick定义一种间隔,20u, 5u, 还可增加3u,4u等, 大的刻度会覆盖小的刻度
- # 20u的间隔,定义大的ticks,并显示ticks_label
- <tick>
- # spacing定义多大间距一个tick
- spacing = 20u
- show_label = yes
- label_size = 20p
- # label的偏移量
- label_offset = 10p
- # %d, %.nf
- format = %d
- </tick>
- # 5u的间隔,定义小的ticks
- <tick>
- spacing = 5u
- color = grey
- size = 10p
- # 定义在哪些染色体显示ticks,哪些区域不显示
- chromosomes=-hs1;-hs2:0-100;-hs3:100-)
- </tick>
- # 30u的间隔,定义中等的ticks
- <tick>
- chromosomes_display_default=no
- spacing = 30u
- color = red
- size = 15p
- chromosomes=hs1
- </tick>
- </ticks>
- ################################################################
- # The remaining content is standard and required. It is imported
- # from default files in the Circos distribution.
- #
- # These should be present in every Circos configuration file and
- # overridden as required. To see the content of these files,
- # look in etc/ in the Circos distribution.
- # 这些都是引用文件,暂时不去管什么意思,后面用到再逐个解释。
- # 但是绘图时这些必须引用。下面会解释下最关键的引用位置。
- <image>
- # Included from Circos distribution.
- <<include etc/image.conf>>
- file* = circos_label_band_tick.png
- </image>
- # RGB/HSV color definitions, color lists, location of fonts, fill patterns.
- # Included from Circos distribution.
- <<include etc/colors_fonts_patterns.conf>>
- # Debugging, I/O an dother system parameters
- # Included from Circos distribution.
- <<include etc/housekeeping.conf>>
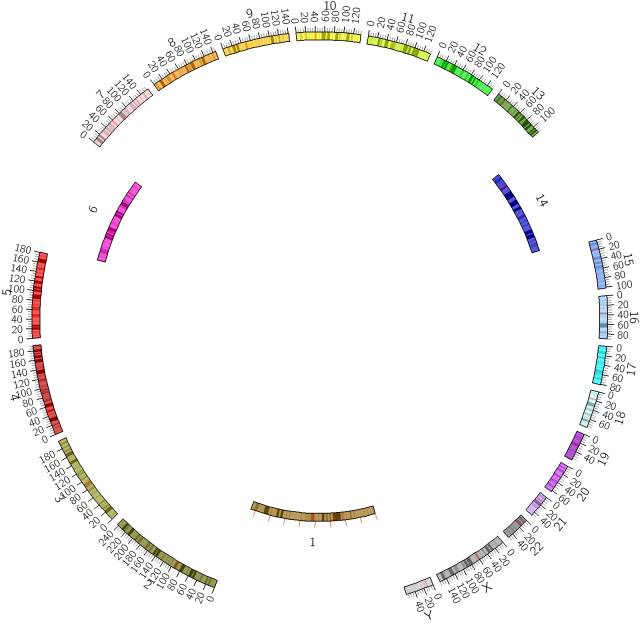
调整染色体位置、大小
在CIRCOS中chromosome表示karyotype文件中定义的染色体的序列结构信息。ideogram是chromosome信息的展示方式。一个染色体可以不展示,也有可能分成多段展示,可以选择图的位置,设置每个染色体的位置、相对大小。
- # karyotype定义染色体的名字、ID、起始位置信息,是绘制图的根本
- karyotype = data/karyotype/karyotype.human.txt
- # 必须的部分,控制染色体信息显示
- # 更多解释见
- # http://circos.ca/documentation/tutorials/reference/ideogram/
- <ideogram>
- # 定义显示的染色体之间的间距,为图形半径的0.5% (r代表radius,半径)
- <spacing>
- default = 0.005r
- </spacing>
- # 染色体区域的绘制位置,默认所有染色体都处于远离圆心同样距离的位置
- # 这里设置的是图形半径的0.9倍的位置
- # 也可以设置绝对像素值
- radius = 0.9r
- # 改变染色体在circos环中内半径大小
- # 染色体区域的宽度,可以是相对图形半径,也可以说绝对像素值
- thickness = 20p
- # 染色体区域填充颜色
- fill = yes
- # 染色体边的颜色和宽度
- stroke_thickness = 2
- stroke_color = black
- # 显示染色体标签名字
- # 更多label的调整见
- # http://circos.ca/documentation/tutorials/ideograms/labels/lesson
- show_label = yes
- label_font = default
- # ideogram外圈再加总半径的0.05
- label_radius = dims(ideogram,radius_outer) + 0.05r
- # 如果想把label标记在圈里可以写,外圈半径+内圈半径除以2
- # label_rdius = (dims(ideogram,radius_outer)+dims(ideogram,radius_inner))/2
- label_size = 24
- # label与圈平行
- label_parallel = yes
- label_case = upper
- # 显示karyotype中定义的细胞遗传学条带
- # fill_bands=yes 表示使用预先定义的颜色
- show_bands = yes
- fill_bands = yes
- band_transparency = 4
- </ideogram>
- # 染色体大小调整
- chromosomes_scale = hs1=0.1r;hs14=0.06r;hs6=0.06r
- # 个别染色体位置调整
- chromosomes_radius = hs14:0.8r;hs6:0.8r;hs1:0.7r
- chromosomes_units = 1000000
- show_ticks = yes
- show_tick_labels = yes
- <ticks>
- # ticks块定义了全局水平的tick的属性
- # 更多解释见http://circos.ca/documentation/tutorials/ticks_and_labels/basics/
- # ticks出现的位置
- radius = dims(ideogram,radius_outer)
- # The label multiplier is the constant used to multiply the tick value to obtain the tick label. For example, if the multiplier is 1e-6, then the tick mark at position 10, 000, 000 will have a label of 10. The multiplier is applied to the raw tick value, regardless of the value of chromosomes_unit.
- # multiplier是用于获得ticks label的,当前ticks对应的染色体位置乘以multiplier就得到ticks label
- # 这个值的设置与chromosomes_units是没有任何关系的
- # 比如当前位置是10,000,000, 乘以multiplier (1e-6)就是10
- multiplier = 1e-6
- # ticks颜色、粗细、大小
- color = black
- thickness = 2p
- size = 15p
- # 默认所有染色体都显示, 放置在ticks块中有效
- chromosomes_display_default=yes
- # 不在任何染色体显示, 放置在ticks块中有效
- #chromosomes_display_default=no
- # 每个tick定义一种间隔,20u, 5u, 还可增加3u,4u等, 大的刻度会覆盖小的刻度
- # 20u的间隔,定义大的ticks,并显示ticks_label
- <tick>
- # spacing定义多大间距一个tick
- spacing = 20u
- show_label = yes
- label_size = 20p
- # label的偏移量
- label_offset = 10p
- # %d, %.nf
- format = %d
- chromosomes=-hs1;-hs14;-hs6
- </tick>
- # 5u的间隔,定义小的ticks
- <tick>
- spacing = 5u
- color = grey
- size = 10p
- # 定义在哪些染色体显示ticks,哪些区域不显示
- chromosomes=-hs1;-hs14;-hs6;-hs2:0-100;-hs3:100-)
- </tick>
- # 30u的间隔,定义中等的ticks
- <tick>
- chromosomes_display_default=no
- spacing = 30u
- color = red
- size = 15p
- chromosomes=hs1
- </tick>
- </ticks>
- ################################################################
- # The remaining content is standard and required. It is imported
- # from default files in the Circos distribution.
- #
- # These should be present in every Circos configuration file and
- # overridden as required. To see the content of these files,
- # look in etc/ in the Circos distribution.
- # 这些都是引用文件,暂时不去管什么意思,后面用到再逐个解释。
- # 但是绘图时这些必须引用。下面会解释下最关键的引用位置。
- <image>
- # Included from Circos distribution.
- <<include etc/image.conf>>
- file* = circos_label_band_tick_chromove.png
- #画图起始的位置,相对于三点钟方向
- angle_offset* = 74
- </image>
- # RGB/HSV color definitions, color lists, location of fonts, fill patterns.
- # Included from Circos distribution.
- <<include etc/colors_fonts_patterns.conf>>
- # Debugging, I/O an dother system parameters
- # Included from Circos distribution.
- <<include etc/housekeeping.conf>>
CIRCOS错误信息
- Dimension [$DIMS->{‘ideogram’}{‘default’}{
' radius_outer'}] isnot definedin expression [dims(ideogram, default, radius_outer) + 0.05r]
radius_outer前面多了个空格
- CONFIGURATION FILE ERROR; Circos could
not findthe configuration file [simple.confy]]
配置文件不存在,或名字、路径写错了
- Config::General: Block ““ has no EndBlock statement (level: 2, chunk 4191)!
区块标签不成对