在使用liger整合单细胞RNA-seq的文章中,我提到liger的数据结构和函数调用不及seurat那么方便和个性化,因此将两者的优势结合起来能够大大便利我们的单细胞数据分析。本文主要介绍以下两种方法:
- 使用SeuratWrappers在seurat中直接调用liger进行降维聚类
- 使用liger内置的函数在liger对象和seurat对象之间进行转换
SeuratWrappers
SeuratWrappers相当于seurat的社区工具,能够使得我们很方便地调用别的包的一些代码和方法来处理seurat对象。SeuratWrappers目前支持Monocle3、glmpca、LIGER、Harmony以及Velocity等13个常用的单细胞转录组相关的包,具体的细节可以参考GitHub。这里不再对代码做任何讲解,因为使用方法和参数与liger基本完全一致,如果有困惑可以参考使用liger整合单细胞RNA-seq。
- ############################################
- ## Project: Liger-learning
- ## Script Purpose: Integrating Seurat objects using LIGER
- ## Data: 2020.11.01
- ## Author: Yiming Sun
- ############################################
- # general setting
- setwd('~/sunym/project/liger_learning/')
- # library
- library(liger)
- library(Seurat)
- library(dplyr)
- library(tidyverse)
- library(viridis)
- library(SeuratData)
- library(SeuratWrappers)
- # 1.Systematic comparative analysis of human PBMC
- data("pbmcsca")
- #pbmcsca is a seurat object
- pbmcsca <- NormalizeData(pbmcsca)
- pbmcsca <- FindVariableFeatures(pbmcsca,selection.method = 'vst',nfeatures = 2000)
- #scale by dfferent methods --> intagrate different methods
- pbmcsca <- ScaleData(pbmcsca,split.by = 'Method',do.center = FALSE)
- pbmcsca <- RunOptimizeALS(pbmcsca,k = 20,lambda = 5,split.by = 'Method',max.iters = 30,thresh = 1e-06)
- pbmcsca <- RunQuantileNorm(pbmcsca,split.by = 'Method',knn_k = 20,quantiles = 50,min_cells = 20,do.center = FALSE,
- max_sample = 1000,refine.knn = TRUE,eps = 0.9)
- #can further cluster the data and find neighbours
- pbmcsca <- FindNeighbors(pbmcsca,reduction = 'iNMF',dims = 1:20)
- pbmcsca <- FindClusters(pbmcsca,resolution = 0.3)
- #dimension reduction and plotting
- pbmcsca <- RunUMAP(pbmcsca,dims = 1:20,reduction = 'iNMF')
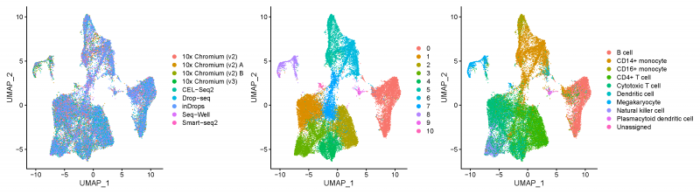
- pdf(file = './res/fig_201101/pbmc_split_by_methods.pdf',width = 18,height = 5)
- DimPlot(pbmcsca,group.by = c('Method','RNA_snn_res.0.3','CellType'),ncol = 3)
- dev.off()
Liger内置函数
Liger包中内置了两个函数ligerToSeurat和seuratToLiger,通常我们用的比较多的是将降维聚类过后的liger对象转换成seurat对象用于做后续的差异表达分析。我们可以简单的来看一下这两个函数的效果。首先先创建一个liger对象。
- ############################################
- ## Project: Liger-learning
- ## Script Purpose: liger and seurat
- ## Data: 2020.11.14
- ## Author: Yiming Sun
- ############################################
- # general setting
- setwd('/data/User/sunym/project/liger_learning/')
- #libarry
- library(liger)
- library(Seurat)
- library(dplyr)
- library(tidyverse)
- library(viridis)
- #######################################
- #liger to seurat
- #######################################
- #load data
- ctrl_dge <- readRDS("./data/PBMC_control.RDS")
- stim_dge <- readRDS("./data/PBMC_interferon-stimulated.RDS")
- #initialize a liger object
- ifnb_liger <- createLiger(list(ctrl = ctrl_dge, stim = stim_dge))
- #explore liger object
- dim([email protected]$ctrl)
- head(colnames([email protected]$ctrl))
- head(rownames([email protected]$ctrl))
- dim([email protected]$stim)
- #normalize data
- ifnb_liger <- normalize(ifnb_liger)
- #select variable gene
- ifnb_liger <- selectGenes(ifnb_liger)
- #scale data but not center
- ifnb_liger <- scaleNotCenter(ifnb_liger)
- #integrate NMF
- ifnb_liger <- optimizeALS(ifnb_liger,k = 20,lambda = 5,max.iters = 30,thresh = 1e-06)
- #Quantile Normalization and Joint Clustering
- ifnb_liger <- quantile_norm(ifnb_liger,knn_k = 20,quantiles = 50,min_cells = 20,do.center = FALSE,
- max_sample = 1000,refine.knn = TRUE,eps = 0.9)
- # you can use louvain cluster to detect and assign cluster
- ifnb_liger <- louvainCluster(ifnb_liger, resolution = 0.25)
- #Visualization and Downstream Analysis
- ifnb_liger <- runUMAP(ifnb_liger, distance = 'cosine', n_neighbors = 30, min_dist = 0.3)
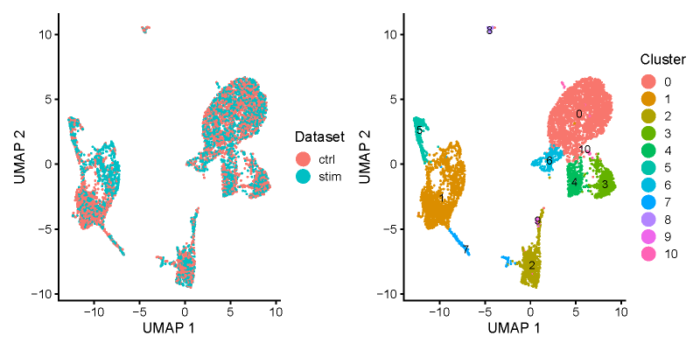
- all.plots <- plotByDatasetAndCluster(ifnb_liger, axis.labels = c('UMAP 1', 'UMAP 2'), return.plots = T)
- pdf(file = './res/fig_201114/plot_by_dataset_and_cluster.pdf',width = 8,height = 4)
- all.plots[[1]] all.plots[[2]]
- dev.off()
- #liger to seurat
- #use nms
- ifnb_seurat <- ligerToSeurat(ifnb_liger,use.liger.genes = TRUE,by.dataset = FALSE,renormalize = TRUE)
- table([email protected])
- head([email protected])
- head(colnames(ifnb_seurat))
可以直接用ligerToSeurat函数进行转换,use.liger.genes参数表示是否保留variable gene的信息,by.dataset参数表示是否在cluster的名字之前加入dataset的名字以作区分,另外默认nms参数为names([email protected]),这个参数会在细胞的barcode之前加入dataset的名称并在orig.ident中标注出数据集的来源。可以看下输出简单理解下。
- > #liger to seurat
- > #use nms
- > ifnb_seurat <- ligerToSeurat(ifnb_liger,use.liger.genes = TRUE,by.dataset = FALSE,renormalize = TRUE)
- Warning: No assay specified, setting assay as RNA by default.
- Warning: No columnames present in cell embeddings, setting to 'iNMF_1:20'
- Warning: No assay specified, setting assay as RNA by default.
- Warning: No columnames present in cell embeddings, setting to 'tSNE_1:2'
- Warning: Feature names cannot have underscores ('_'), replacing with dashes ('-')
- Performing log-normalization
- 0% 10 20 30 40 50 60 70 80 90 100%
- [----|----|----|----|----|----|----|----|----|----|
- **************************************************|
- > table([email protected])
- 1 0 2 9 4 3 6 7 5 8 10
- 1500 2309 609 37 384 444 238 140 251 56 32
- > head([email protected])
- orig.ident nCount_RNA nFeature_RNA
- ctrl_ctrlTCAGCGCTGGTCAT-1 ctrl 2232 815
- ctrl_ctrlTTATGGCTTCATTC-1 ctrl 2466 760
- ctrl_ctrlACCCACTGCTTAGG-1 ctrl 1085 452
- ctrl_ctrlATGGGTACCCCGTT-1 ctrl 3242 925
- ctrl_ctrlTGACTGGACAGTCA-1 ctrl 635 333
- ctrl_ctrlGTGTAGTGGTTGTG-1 ctrl 1462 549
- > head(colnames(ifnb_seurat))
- [1] "ctrl_ctrlTCAGCGCTGGTCAT-1" "ctrl_ctrlTTATGGCTTCATTC-1"
- [3] "ctrl_ctrlACCCACTGCTTAGG-1" "ctrl_ctrlATGGGTACCCCGTT-1"
- [5] "ctrl_ctrlTGACTGGACAGTCA-1" "ctrl_ctrlGTGTAGTGGTTGTG-1"
如果令nms = NULL。
- > #not use nms
- > ifnb_seurat <- ligerToSeurat(ifnb_liger,nms = NULL,use.liger.genes = TRUE,by.dataset = FALSE,renormalize = TRUE)
- Warning: No assay specified, setting assay as RNA by default.
- Warning: No columnames present in cell embeddings, setting to 'iNMF_1:20'
- Warning: No assay specified, setting assay as RNA by default.
- Warning: No columnames present in cell embeddings, setting to 'tSNE_1:2'
- Warning: Feature names cannot have underscores ('_'), replacing with dashes ('-')
- Performing log-normalization
- 0% 10 20 30 40 50 60 70 80 90 100%
- [----|----|----|----|----|----|----|----|----|----|
- **************************************************|
- > table([email protected])
- 1 0 2 9 4 3 6 7 5 8 10
- 1500 2309 609 37 384 444 238 140 251 56 32
- > head([email protected])
- orig.ident nCount_RNA nFeature_RNA
- ctrlTCAGCGCTGGTCAT-1 SeuratProject 2232 815
- ctrlTTATGGCTTCATTC-1 SeuratProject 2466 760
- ctrlACCCACTGCTTAGG-1 SeuratProject 1085 452
- ctrlATGGGTACCCCGTT-1 SeuratProject 3242 925
- ctrlTGACTGGACAGTCA-1 SeuratProject 635 333
- ctrlGTGTAGTGGTTGTG-1 SeuratProject 1462 549
- > head(colnames(ifnb_seurat))
- [1] "ctrlTCAGCGCTGGTCAT-1" "ctrlTTATGGCTTCATTC-1"
- [3] "ctrlACCCACTGCTTAGG-1" "ctrlATGGGTACCCCGTT-1"
- [5] "ctrlTGACTGGACAGTCA-1" "ctrlGTGTAGTGGTTGTG-1"
使用seurat的函数做个性化的差异表达分析,参考文章Seurat进行单细胞RNA-seq聚类分析。
- #use liger cluster as cell type and do the DE analysis
- new.cluster.ids <- c("a", "b", "c", "d", "e", "f", "g", "h", "i", "j", "k")
- names(new.cluster.ids) <- levels([email protected])
- ifnb_seurat <- RenameIdents(ifnb_seurat, new.cluster.ids)
- ifnb_seurat$cell_type <- [email protected]
- Idents(ifnb_seurat) <- 'cell_type'
- all.marker <- FindAllMarkers(ifnb_seurat,only.pos = TRUE,min.pct = 0.25,logfc.threshold = 0.25)
- cluster_a_vs_b_marker <- FindMarkers(ifnb_seurat,group.by = 'cell_type',ident.1 = 'a',ident.2 = 'b',only.pos = TRUE)
手动导出liger中的降维图并导入seurat。
- #get the tsne manually
- tsne.obj <- CreateDimReducObject(embeddings = [email protected],key = 'testUMAP_',global = TRUE)
- ifnb_seurat[['tsne']] <- tsne.obj
- pdf(file = './res/fig_201114/dimplot_tsetUMAP.pdf',width = 9,height = 5)
- DimPlot(ifnb_seurat,group.by = 'cell_type')
- dev.off()
注意tsne.obj的barcode要与seurat中的barcode相对应,因此可以将nms设为NULL或者为tsne.obj手动加上dataset的标签。
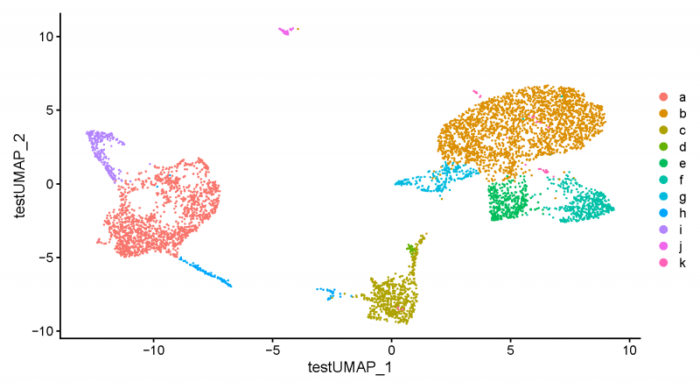
可以看到key参数中的内容被成功导入进去了。最后我们也可以将seurat对象转换为liger对象。
- > ###################################################
- > #seurat to liger
- > ###################################################
- > ifnb_liger <- seuratToLiger(ifnb_seurat,combined.seurat = TRUE,meta.var = 'orig.ident',renormalize = TRUE)
- > head([email protected])
- ctrlTCAGCGCTGGTCAT-1 ctrlTTATGGCTTCATTC-1 ctrlACCCACTGCTTAGG-1
- a a b
- ctrlATGGGTACCCCGTT-1 ctrlTGACTGGACAGTCA-1 ctrlGTGTAGTGGTTGTG-1
- a c b
- Levels: a b c d e f g h i j k
- > head([email protected])
- tSNE_1 tSNE_2
- ctrlTCAGCGCTGGTCAT-1 -9.5633178 -1.5025842
- ctrlTTATGGCTTCATTC-1 -7.4026990 -0.5618219
- ctrlACCCACTGCTTAGG-1 3.7179575 4.9839707
- ctrlATGGGTACCCCGTT-1 -11.0730367 -4.6024990
- ctrlTGACTGGACAGTCA-1 0.5273923 -8.4171533
- ctrlGTGTAGTGGTTGTG-1 8.8018236 5.2826267
写在最后
这部分的内容比较枯燥,主要是我自己探索了一下seurat和liger的数据结构以及他们之间如何进行相互转换,想在这里记录一下以免自己忘了。


