在我们分析单细胞数据的时候,需要想象力的一点就是要理解数据结构。平时我们都是如何看数据结构的呢?
- library(Seurat)
- library(tidyverse)
- pbmc<-CreateSeuratObject(pbmc_small@assays$RNA@counts)
- pbmc%>% NormalizeData() %>% FindVariableFeatures() %>%
- ScaleData() %>% RunPCA() %>% FindNeighbors() %>% RunUMAP(1:10) %>%
- FindClusters(dims=1:0)-> pbmc
- pbmc
- An object of class Seurat
- 230 features across 80 samples within 1 assay
- Active assay: RNA (230 features)
- 2 dimensional reductions calculated: pca, umap
在R里面我们用的是str(...),如:
- str(pbmc)
- Formal class 'Seurat' [package "Seurat"] with 13 slots
- ..@ assays :List of 1
- .. ..$ RNA:Formal class 'Assay' [package "Seurat"] with 8 slots
- .. .. .. ..@ counts :Formal class 'dgCMatrix' [package "Matrix"] with 6 slots
- .. .. .. .. .. ..@ i : int [1:4456] 1 5 8 11 22 30 33 34 36 38 ...
- .. .. .. .. .. ..@ p : int [1:81] 0 47 99 149 205 258 306 342 387 423 ...
- .. .. .. .. .. ..@ Dim : int [1:2] 230 80
- .. .. .. .. .. ..@ Dimnames:List of 2
- .. .. .. .. .. .. ..$ : chr [1:230] "MS4A1" "CD79B" "CD79A" "HLA-DRA" ...
- .. .. .. .. .. .. ..$ : chr [1:80] "ATGCCAGAACGACT" "CATGGCCTGTGCAT" "GAACCTGATGAACC" "TGACTGGATTCTCA" ...
- .. .. .. .. .. ..@ x : num [1:4456] 1 1 3 1 1 4 1 5 1 1 ...
- .. .. .. .. .. ..@ factors : list()
- .. .. .. ..@ data :Formal class 'dgCMatrix' [package "Matrix"] with 6 slots
- .. .. .. .. .. ..@ i : int [1:4456] 1 5 8 11 22 30 33 34 36 38 ...
- .. .. .. .. .. ..@ p : int [1:81] 0 47 99 149 205 258 306 342 387 423 ...
- .. .. .. .. .. ..@ Dim : int [1:2] 230 80
- .. .. .. .. .. ..@ Dimnames:List of 2
- .. .. .. .. .. .. ..$ : chr [1:230] "MS4A1" "CD79B" "CD79A" "HLA-DRA" ...
- .. .. .. .. .. .. ..$ : chr [1:80] "ATGCCAGAACGACT" "CATGGCCTGTGCAT" "GAACCTGATGAACC" "TGACTGGATTCTCA" ...
- .. .. .. .. .. ..@ x : num [1:4456] 4.97 4.97 6.06 4.97 4.97 ...
- .. .. .. .. .. ..@ factors : list()
- .. .. .. ..@ scale.data : num [1:230, 1:80] -0.409 1.64 -0.428 -1.375 -0.329 ...
- .. .. .. .. ..- attr(*, "dimnames")=List of 2
- .. .. .. .. .. ..$ : chr [1:230] "MS4A1" "CD79B" "CD79A" "HLA-DRA" ...
- .. .. .. .. .. ..$ : chr [1:80] "ATGCCAGAACGACT" "CATGGCCTGTGCAT" "GAACCTGATGAACC" "TGACTGGATTCTCA" ...
- .. .. .. ..@ key : chr "rna_"
- .. .. .. ..@ assay.orig : NULL
- .. .. .. ..@ var.features : chr [1:230] "PPBP" "IGLL5" "VDAC3" "CD1C" ...
- .. .. .. ..@ meta.features:'data.frame': 230 obs. of 5 variables:
- .. .. .. .. ..$ vst.mean : num [1:230] 0.388 0.6 0.7 13.425 0.3 ...
- .. .. .. .. ..$ vst.variance : num [1:230] 1.025 1.281 4.365 725.463 0.871 ...
- .. .. .. .. ..$ vst.variance.expected : num [1:230] 1.141 2.664 4.029 745.145 0.642 ...
- .. .. .. .. ..$ vst.variance.standardized: num [1:230] 0.898 0.481 1.083 0.974 1.356 ...
- .. .. .. .. ..$ vst.variable : logi [1:230] TRUE TRUE TRUE TRUE TRUE TRUE ...
- .. .. .. ..@ misc : NULL
- ..@ meta.data :'data.frame': 80 obs. of 5 variables:
- .. ..$ orig.ident : Factor w/ 1 level "SeuratProject": 1 1 1 1 1 1 1 1 1 1 ...
- .. ..$ nCount_RNA : num [1:80] 70 85 87 127 173 70 64 72 52 100 ...
- .. ..$ nFeature_RNA : int [1:80] 47 52 50 56 53 48 36 45 36 41 ...
- .. ..$ RNA_snn_res.0.8: Factor w/ 3 levels "0","1","2": 2 2 2 2 2 2 2 2 2 2 ...
- .. ..$ seurat_clusters: Factor w/ 3 levels "0","1","2": 2 2 2 2 2 2 2 2 2 2 ...
- ..@ active.assay: chr "RNA"
- ..@ active.ident: Factor w/ 3 levels "0","1","2": 2 2 2 2 2 2 2 2 2 2 ...
- .. ..- attr(*, "names")= chr [1:80] "ATGCCAGAACGACT" "CATGGCCTGTGCAT" "GAACCTGATGAACC" "TGACTGGATTCTCA" ...
- ..@ graphs :List of 2
- .. ..$ RNA_nn :Formal class 'Graph' [package "Seurat"] with 7 slots
- .. .. .. ..@ assay.used: chr "RNA"
- .. .. .. ..@ i : int [1:1600] 0 1 2 3 4 5 6 7 8 9 ...
- .. .. .. ..@ p : int [1:81] 0 10 17 40 57 101 124 141 153 178 ...
- .. .. .. ..@ Dim : int [1:2] 80 80
- .. .. .. ..@ Dimnames :List of 2
- .. .. .. .. ..$ : chr [1:80] "ATGCCAGAACGACT" "CATGGCCTGTGCAT" "GAACCTGATGAACC" "TGACTGGATTCTCA" ...
- .. .. .. .. ..$ : chr [1:80] "ATGCCAGAACGACT" "CATGGCCTGTGCAT" "GAACCTGATGAACC" "TGACTGGATTCTCA" ...
- .. .. .. ..@ x : num [1:1600] 1 1 1 1 1 1 1 1 1 1 ...
- .. .. .. ..@ factors : list()
- .. ..$ RNA_snn:Formal class 'Graph' [package "Seurat"] with 7 slots
- .. .. .. ..@ assay.used: chr "RNA"
- .. .. .. ..@ i : int [1:4174] 0 1 2 3 4 5 6 7 8 9 ...
- .. .. .. ..@ p : int [1:81] 0 68 132 181 230 277 326 375 424 487 ...
- .. .. .. ..@ Dim : int [1:2] 80 80
- .. .. .. ..@ Dimnames :List of 2
- .. .. .. .. ..$ : chr [1:80] "ATGCCAGAACGACT" "CATGGCCTGTGCAT" "GAACCTGATGAACC" "TGACTGGATTCTCA" ...
- .. .. .. .. ..$ : chr [1:80] "ATGCCAGAACGACT" "CATGGCCTGTGCAT" "GAACCTGATGAACC" "TGACTGGATTCTCA" ...
- .. .. .. ..@ x : num [1:4174] 1 0.6 0.6 0.6 0.538 ...
- .. .. .. ..@ factors : list()
- ..@ neighbors : list()
- ..@ reductions :List of 2
- .. ..$ pca :Formal class 'DimReduc' [package "Seurat"] with 9 slots
- .. .. .. ..@ cell.embeddings : num [1:80, 1:50] 3.12 3.56 2.4 3.43 2.78 ...
- .. .. .. .. ..- attr(*, "dimnames")=List of 2
- .. .. .. .. .. ..$ : chr [1:80] "ATGCCAGAACGACT" "CATGGCCTGTGCAT" "GAACCTGATGAACC" "TGACTGGATTCTCA" ...
- .. .. .. .. .. ..$ : chr [1:50] "PC_1" "PC_2" "PC_3" "PC_4" ...
- .. .. .. ..@ feature.loadings : num [1:230, 1:50] 0.05711 0.00738 0.03005 -0.04766 0.05598 ...
- .. .. .. .. ..- attr(*, "dimnames")=List of 2
- .. .. .. .. .. ..$ : chr [1:230] "PPBP" "IGLL5" "VDAC3" "CD1C" ...
- .. .. .. .. .. ..$ : chr [1:50] "PC_1" "PC_2" "PC_3" "PC_4" ...
- .. .. .. ..@ feature.loadings.projected: num[0 , 0 ]
- .. .. .. ..@ assay.used : chr "RNA"
- .. .. .. ..@ global : logi FALSE
- .. .. .. ..@ stdev : num [1:50] 5.75 5.21 4.32 3.62 2.77 ...
- .. .. .. ..@ key : chr "PC_"
- .. .. .. ..@ jackstraw :Formal class 'JackStrawData' [package "Seurat"] with 4 slots
- .. .. .. .. .. ..@ empirical.p.values : num[0 , 0 ]
- .. .. .. .. .. ..@ fake.reduction.scores : num[0 , 0 ]
- .. .. .. .. .. ..@ empirical.p.values.full: num[0 , 0 ]
- .. .. .. .. .. ..@ overall.p.values : num[0 , 0 ]
- .. .. .. ..@ misc :List of 1
- .. .. .. .. ..$ total.variance: num 230
- .. ..$ umap:Formal class 'DimReduc' [package "Seurat"] with 9 slots
- .. .. .. ..@ cell.embeddings : num [1:80, 1:2] 5.07 5.31 4.72 5.06 5.45 ...
- .. .. .. .. ..- attr(*, "scaled:center")= num [1:2] 1.78 -8.75
- .. .. .. .. ..- attr(*, "dimnames")=List of 2
- .. .. .. .. .. ..$ : chr [1:80] "ATGCCAGAACGACT" "CATGGCCTGTGCAT" "GAACCTGATGAACC" "TGACTGGATTCTCA" ...
- .. .. .. .. .. ..$ : chr [1:2] "UMAP_1" "UMAP_2"
- .. .. .. ..@ feature.loadings : num[0 , 0 ]
- .. .. .. ..@ feature.loadings.projected: num[0 , 0 ]
- .. .. .. ..@ assay.used : chr "RNA"
- .. .. .. ..@ global : logi TRUE
- .. .. .. ..@ stdev : num(0)
- .. .. .. ..@ key : chr "UMAP_"
- .. .. .. ..@ jackstraw :Formal class 'JackStrawData' [package "Seurat"] with 4 slots
- .. .. .. .. .. ..@ empirical.p.values : num[0 , 0 ]
- .. .. .. .. .. ..@ fake.reduction.scores : num[0 , 0 ]
- .. .. .. .. .. ..@ empirical.p.values.full: num[0 , 0 ]
- .. .. .. .. .. ..@ overall.p.values : num[0 , 0 ]
- .. .. .. ..@ misc : list()
- ..@ images : list()
- ..@ project.name: chr "SeuratProject"
- ..@ misc : list()
- ..@ version :Classes 'package_version', 'numeric_version' hidden list of 1
- .. ..$ : int [1:3] 3 1 2
- ..@ commands :List of 7
- .. ..$ NormalizeData.RNA :Formal class 'SeuratCommand' [package "Seurat"] with 5 slots
- .. .. .. ..@ name : chr "NormalizeData.RNA"
- .. .. .. ..@ time.stamp : POSIXct[1:1], format: "2020-06-01 22:43:27"
- .. .. .. ..@ assay.used : chr "RNA"
- .. .. .. ..@ call.string: chr "NormalizeData(.)"
- .. .. .. ..@ params :List of 5
- .. .. .. .. ..$ assay : chr "RNA"
- .. .. .. .. ..$ normalization.method: chr "LogNormalize"
- .. .. .. .. ..$ scale.factor : num 10000
- .. .. .. .. ..$ margin : num 1
- .. .. .. .. ..$ verbose : logi TRUE
- .. ..$ FindVariableFeatures.RNA:Formal class 'SeuratCommand' [package "Seurat"] with 5 slots
- .. .. .. ..@ name : chr "FindVariableFeatures.RNA"
- .. .. .. ..@ time.stamp : POSIXct[1:1], format: "2020-06-01 22:43:28"
- .. .. .. ..@ assay.used : chr "RNA"
- .. .. .. ..@ call.string: chr "FindVariableFeatures(.)"
- .. .. .. ..@ params :List of 12
- .. .. .. .. ..$ assay : chr "RNA"
- .. .. .. .. ..$ selection.method : chr "vst"
- .. .. .. .. ..$ loess.span : num 0.3
- .. .. .. .. ..$ clip.max : chr "auto"
- .. .. .. .. ..$ mean.function :function (mat, display_progress)
- .. .. .. .. ..$ dispersion.function:function (mat, display_progress)
- .. .. .. .. ..$ num.bin : num 20
- .. .. .. .. ..$ binning.method : chr "equal_width"
- .. .. .. .. ..$ nfeatures : num 2000
- .. .. .. .. ..$ mean.cutoff : num [1:2] 0.1 8
- .. .. .. .. ..$ dispersion.cutoff : num [1:2] 1 Inf
- .. .. .. .. ..$ verbose : logi TRUE
- .. ..$ ScaleData.RNA :Formal class 'SeuratCommand' [package "Seurat"] with 5 slots
- .. .. .. ..@ name : chr "ScaleData.RNA"
- .. .. .. ..@ time.stamp : POSIXct[1:1], format: "2020-06-01 22:43:28"
- .. .. .. ..@ assay.used : chr "RNA"
- .. .. .. ..@ call.string: chr "ScaleData(.)"
- .. .. .. ..@ params :List of 10
- .. .. .. .. ..$ features : chr [1:230] "PPBP" "IGLL5" "VDAC3" "CD1C" ...
- .. .. .. .. ..$ assay : chr "RNA"
- .. .. .. .. ..$ model.use : chr "linear"
- .. .. .. .. ..$ use.umi : logi FALSE
- .. .. .. .. ..$ do.scale : logi TRUE
- .. .. .. .. ..$ do.center : logi TRUE
- .. .. .. .. ..$ scale.max : num 10
- .. .. .. .. ..$ block.size : num 1000
- .. .. .. .. ..$ min.cells.to.block: num 80
- .. .. .. .. ..$ verbose : logi TRUE
- .. ..$ RunPCA.RNA :Formal class 'SeuratCommand' [package "Seurat"] with 5 slots
- .. .. .. ..@ name : chr "RunPCA.RNA"
- .. .. .. ..@ time.stamp : POSIXct[1:1], format: "2020-06-01 22:43:29"
- .. .. .. ..@ assay.used : chr "RNA"
- .. .. .. ..@ call.string: chr "RunPCA(.)"
- .. .. .. ..@ params :List of 10
- .. .. .. .. ..$ assay : chr "RNA"
- .. .. .. .. ..$ npcs : num 50
- .. .. .. .. ..$ rev.pca : logi FALSE
- .. .. .. .. ..$ weight.by.var : logi TRUE
- .. .. .. .. ..$ verbose : logi TRUE
- .. .. .. .. ..$ ndims.print : int [1:5] 1 2 3 4 5
- .. .. .. .. ..$ nfeatures.print: num 30
- .. .. .. .. ..$ reduction.name : chr "pca"
- .. .. .. .. ..$ reduction.key : chr "PC_"
- .. .. .. .. ..$ seed.use : num 42
- .. ..$ FindNeighbors.RNA.pca :Formal class 'SeuratCommand' [package "Seurat"] with 5 slots
- .. .. .. ..@ name : chr "FindNeighbors.RNA.pca"
- .. .. .. ..@ time.stamp : POSIXct[1:1], format: "2020-06-01 22:43:29"
- .. .. .. ..@ assay.used : chr "RNA"
- .. .. .. ..@ call.string: chr "FindNeighbors(.)"
- .. .. .. ..@ params :List of 13
- .. .. .. .. ..$ reduction : chr "pca"
- .. .. .. .. ..$ dims : int [1:10] 1 2 3 4 5 6 7 8 9 10
- .. .. .. .. ..$ assay : chr "RNA"
- .. .. .. .. ..$ k.param : num 20
- .. .. .. .. ..$ compute.SNN : logi TRUE
- .. .. .. .. ..$ prune.SNN : num 0.0667
- .. .. .. .. ..$ nn.method : chr "rann"
- .. .. .. .. ..$ annoy.metric: chr "euclidean"
- .. .. .. .. ..$ nn.eps : num 0
- .. .. .. .. ..$ verbose : logi TRUE
- .. .. .. .. ..$ force.recalc: logi FALSE
- .. .. .. .. ..$ do.plot : logi FALSE
- .. .. .. .. ..$ graph.name : chr [1:2] "RNA_nn" "RNA_snn"
- .. ..$ RunUMAP.RNA.pca :Formal class 'SeuratCommand' [package "Seurat"] with 5 slots
- .. .. .. ..@ name : chr "RunUMAP.RNA.pca"
- .. .. .. ..@ time.stamp : POSIXct[1:1], format: "2020-06-01 22:43:33"
- .. .. .. ..@ assay.used : chr "RNA"
- .. .. .. ..@ call.string: chr "RunUMAP(., 1:10)"
- .. .. .. ..@ params :List of 20
- .. .. .. .. ..$ dims : int [1:10] 1 2 3 4 5 6 7 8 9 10
- .. .. .. .. ..$ reduction : chr "pca"
- .. .. .. .. ..$ assay : chr "RNA"
- .. .. .. .. ..$ umap.method : chr "uwot"
- .. .. .. .. ..$ n.neighbors : int 30
- .. .. .. .. ..$ n.components : int 2
- .. .. .. .. ..$ metric : chr "cosine"
- .. .. .. .. ..$ learning.rate : num 1
- .. .. .. .. ..$ min.dist : num 0.3
- .. .. .. .. ..$ spread : num 1
- .. .. .. .. ..$ set.op.mix.ratio : num 1
- .. .. .. .. ..$ local.connectivity : int 1
- .. .. .. .. ..$ repulsion.strength : num 1
- .. .. .. .. ..$ negative.sample.rate: int 5
- .. .. .. .. ..$ uwot.sgd : logi FALSE
- .. .. .. .. ..$ seed.use : int 42
- .. .. .. .. ..$ angular.rp.forest : logi FALSE
- .. .. .. .. ..$ verbose : logi TRUE
- .. .. .. .. ..$ reduction.name : chr "umap"
- .. .. .. .. ..$ reduction.key : chr "UMAP_"
- .. ..$ FindClusters :Formal class 'SeuratCommand' [package "Seurat"] with 5 slots
- .. .. .. ..@ name : chr "FindClusters"
- .. .. .. ..@ time.stamp : POSIXct[1:1], format: "2020-06-01 22:43:33"
- .. .. .. ..@ assay.used : chr "RNA"
- .. .. .. ..@ call.string: chr "FindClusters(., dims = 1:0)"
- .. .. .. ..@ params :List of 10
- .. .. .. .. ..$ graph.name : chr "RNA_snn"
- .. .. .. .. ..$ modularity.fxn : num 1
- .. .. .. .. ..$ resolution : num 0.8
- .. .. .. .. ..$ method : chr "matrix"
- .. .. .. .. ..$ algorithm : num 1
- .. .. .. .. ..$ n.start : num 10
- .. .. .. .. ..$ n.iter : num 10
- .. .. .. .. ..$ random.seed : num 0
- .. .. .. .. ..$ group.singletons: logi TRUE
- .. .. .. .. ..$ verbose : logi TRUE
- ..@ tools : list()
别说看了,拉鼠标手都能拉疼。那么我们能不能基于str(pbmc)的结果做一个思维导图呢?就像这样:
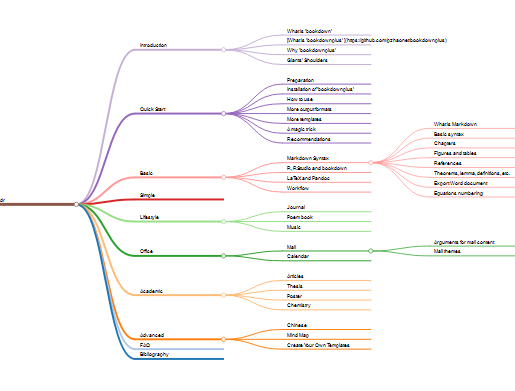
如果能够这样查看,那不是美滋滋的吗?
需求有了,就差行动了,我们来找代码:
- library(mindr)
- (out <- capture.output(str(pbmc)))
- out2 <- paste(out, collapse="n")
- mm(gsub("\.\.@","# ",gsub("\.\. ","#",out2)),type ="text",root= "Seurat")
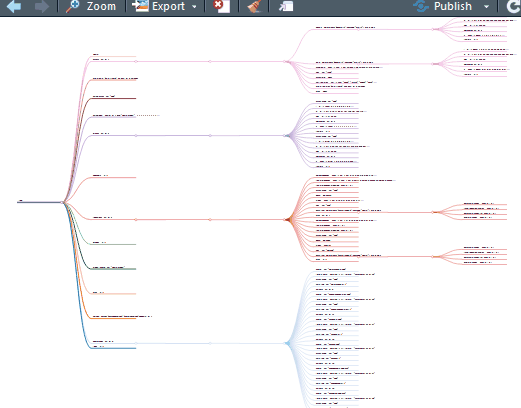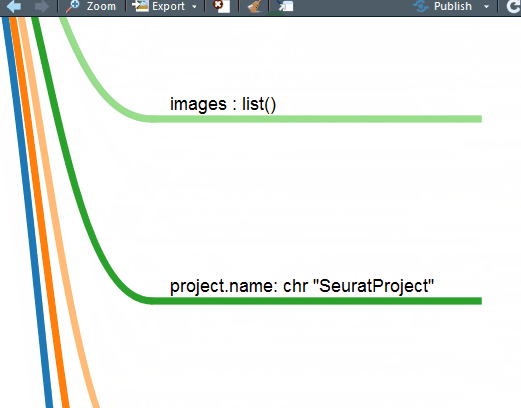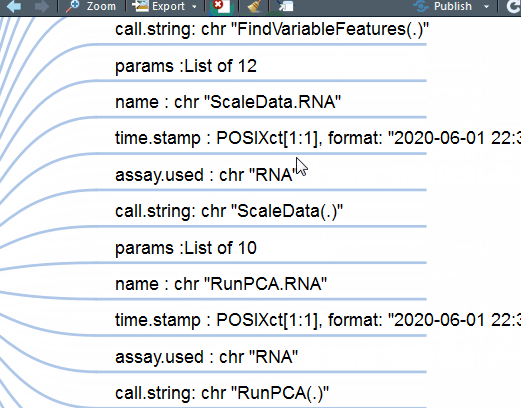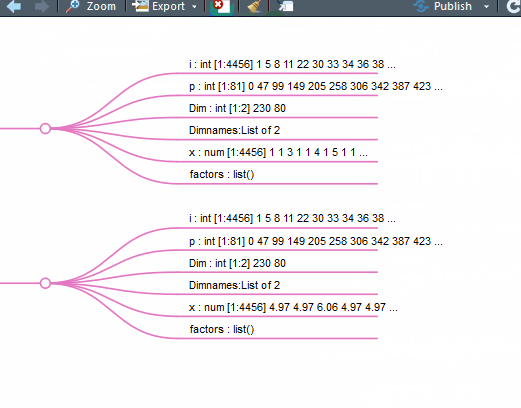
这下好了,你对单细胞Seurat数据对象做了什么一目了然。


